About This File
Here is the Hooovahh Array VIMs. This initial release contains VIMs for manipulating array data, which are intended to replace OpenG functionality, but with the added benefit of data type propagation, and increased performance using newer array manipulation techniques. In later versions other Array manipulation functions were added moving all the OpenG stuff to their own palette.
Version 2.0 changed the suffix naming standard. Updating may mean replacing calls to the new versions since the name on disk has changed. This was for consistency and I'm sorry for breaking compatibility. The added type defs in 2.0 may break compatibility too but these help avoid code breaking bugs since VIMs allowed any data type previously.
Most of the OpenG functions are unchanged, but a few use the newer conditional and concatenating tunnels. And a few functions have added performance based on other inputs. For instance the Delete Array Elements can operate in a more efficient way if the input indexes are already sorted. The Filter 1D array can also be more efficient if the input is known to not contain any duplicates.
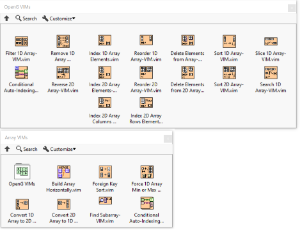
Because these packages contain VIMs, they require LabVIEW 2017 or newer. Having these functions be VIMs mean all functions work with various array data types. Included functions are:
- Conditional Auto-Indexing Tunnel
- Delete Elements from (1D or 2D) Array
- Filter 1D Array
- Index (1D or 2D) Array, Scalar, Row, Column
- Remove Duplicates from 1D Array
- Reorder (1D or 2D) Array
- Reverse 1D Array
- Slice 1D Array
- Sort (1D or 2D) Array
- Convert 1D to 2D
- Convert 2D to 1D
- Find Subarray
- Force Array Min/Max Size
- Foreign Key Sort
What's New in Version 2.0.0 See changelog
Released
- Updated to LabVIEW 2018
- Removed file suffix on installed files
- Fix with delete elements from 2D Array
- Added Index 2D array Row/Column
- Added type def enums for various VIs